 |
SLIMS |
 |
Inventory |
 |
Experiments |
|
|
Compounds
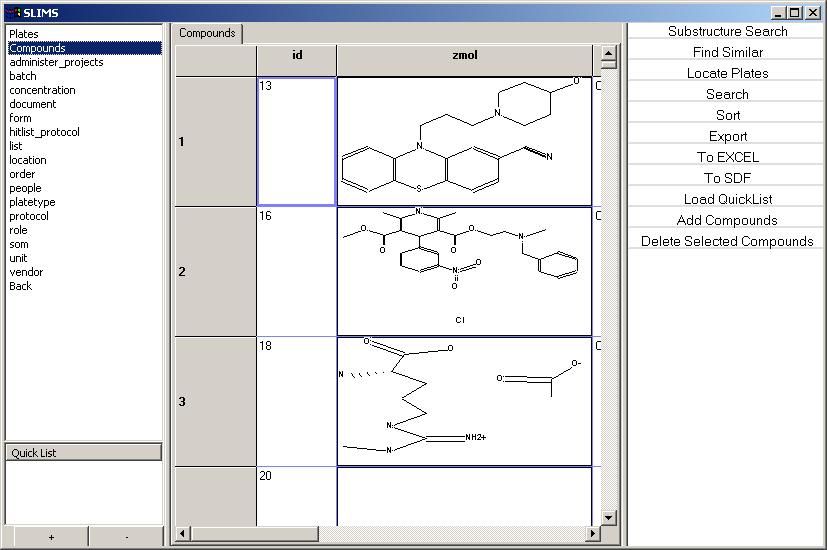
The compound structures are displayed in the view along with the
following properties:
- smiles - the smiles string for the
compound
- name - the name of the compound, either user
supplied or the drug name
- vendor - the vendor of the compound, this comes
from the vender table which is a controlled
vocabulary
- library - this is a user supplied tag to annotate
compounds purchased at different times
- catno - the vendor's catalogue number of the
compound
- mw_total - molecular weight of the compound
- mw_active - the molecular weight of the smallest
fragment of the compound
- num_atoms - the number of atoms in the compound
- num_bonds - the number of bonds in the compound
The compound view has the following commands available:
- Substructure
Search - Search the current compound view for a
substructure. This command opens up a new dialog box where a
substructure can be entered. If a row is selected, then the view
is opened with the substructure from the current row. Exiting
this dialog starts the substructure search.
- Find
Similar - Sorts the compounds based on their similarity to the
currently selected compound.
- Locate
Plates - Display all the plates where the currently selected
compounds are found.
- Search
- Search for properties in the view
- Sort
- Sort the view
- Export
- Export the selected view to a csv file. This doesn't
save images.
- Export
To Excel - Export the selected view to Excel. Excel must
be installed on your computer. This really should only be used
for relatively small views.
- To
SDF - Export the current view as a MDL Structured Data
File. This is a standard format for transporting chemical
structures and their related data fields.
- Load
QuickList - Join the current view with the given
quicklist. Used from the compound view, this shows all
compounds stored in the quicklist.
- Add Compounds - Add Compounds to the
database. This is also how batches are added to the database.
|